Orchid: Revolutionizing Deep Learning with Data-Dependent Convolutions for Scalable Sequence Modeling
In the fast-paced world of deep learning, researchers are constantly striving to develop more efficient and effective models for tasks such as natural language processing, image analysis, and genomics. One of the key challenges in these fields is finding a balance between computational efficiency and expressive power. Traditional attention mechanisms have been groundbreaking in their ability to handle sequence modeling tasks, but they often suffer from quadratic complexity, making them less suitable for handling long-context tasks.
To address this challenge, researchers at the University of Waterloo have introduced a groundbreaking deep learning architecture called Orchid. Orchid leverages a data-dependent convolution mechanism to revolutionize sequence modeling and overcome the limitations of traditional attention-based models. This innovative approach not only improves computational efficiency but also maintains robust expressiveness, making Orchid a game-changer in the field of deep learning.
The Challenge of Sequence Modeling
Sequence modeling is widely used in various domains, including natural language processing, genomics, and image analysis. It involves analyzing and understanding the relationships and dependencies between elements in a sequence of data. Traditional attention mechanisms, such as those used in models like BERT and Vision Transformers, have been instrumental in advancing sequence modeling tasks. However, their computational complexity scales quadratically with sequence length, posing a significant bottleneck when dealing with long-context tasks.
The ever-growing need to process larger and more complex datasets has motivated researchers to find more efficient and scalable solutions. Several approaches, such as sparsifying attention matrices or employing low-rank approximations, have been proposed to reduce the computational burden of attention mechanisms. While these techniques improve efficiency to some extent, they often struggle to strike the perfect balance between computational complexity and expressive power.
Orchid: A New Architectural Innovation
Orchid, pioneered by researchers at the University of Waterloo, introduces a novel data-dependent convolution mechanism to tackle the challenges of sequence modeling. By integrating this mechanism into deep learning architectures, Orchid achieves both computational efficiency and robust expressiveness.
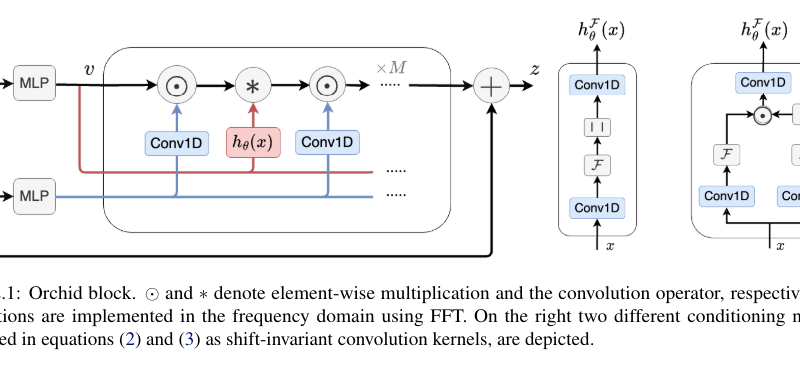
At the core of Orchid lies its data-dependent convolution layer, which dynamically adjusts its kernel based on the input data using a conditioning neural network. This adaptive convolution mechanism allows Orchid to efficiently filter long sequences, enabling scalability with quasi-linear complexity. The conditioning network ensures that the kernel adapts to the input data, enhancing Orchid’s ability to capture long-range dependencies while maintaining computational efficiency.
With the incorporation of gating operations, Orchid achieves high expressivity while still offering quasi-linear scalability, with a complexity of O(LlogL). This means that Orchid can handle sequence lengths well beyond the limitations of dense attention layers, making it suitable for processing increasingly large and complex datasets.
Performance Benchmarks
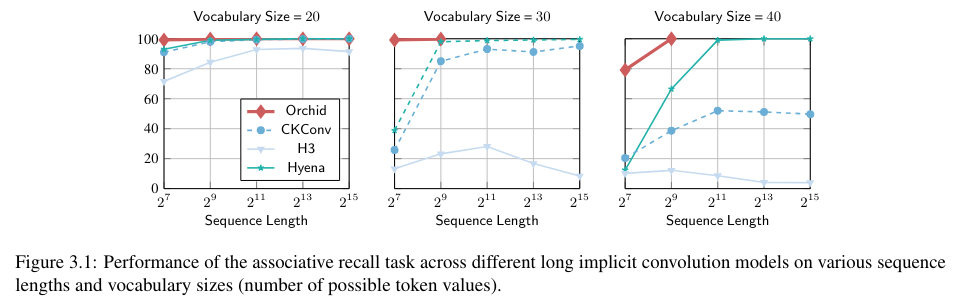
Comparisons between Orchid and traditional attention-based models demonstrate the superior performance of Orchid in various sequence modeling tasks. For instance, on the Associative Recall task, Orchid consistently achieves accuracy rates above 99% for sequences up to 131K. In comparison, the BERT-base model has 30% more parameters but achieves a 1.0-point improvement in the GLUE score. Similarly, Orchid-BERT-large surpasses BERT-large in GLUE performance while reducing parameter counts by 25%.
These performance benchmarks highlight Orchid’s potential as a versatile model for handling increasingly large and complex datasets. By effectively addressing the computational complexity limitations of traditional attention mechanisms, Orchid offers a transformative approach to sequence modeling in deep learning.
The Future of Deep Learning with Orchid
As researchers continue to explore the potential of deep learning in various domains, efficiency and scalability remain crucial factors. Orchid’s innovative approach to sequence modeling opens up new possibilities for developing more efficient and effective deep learning models. By leveraging data-dependent convolutions, Orchid provides a powerful tool for handling long-context tasks and processing large datasets.
The introduction of Orchid has a significant impact on the field of deep learning, offering researchers and practitioners a scalable and expressive solution for sequence modeling. As the capabilities of Orchid continue to evolve, it is expected to drive further advancements in natural language processing, genomics, image analysis, and other related domains.
In conclusion, Orchid, the deep learning architecture developed by researchers at the University of Waterloo, revolutionizes sequence modeling with its data-dependent convolution mechanism. By dynamically adjusting its kernel based on input data, Orchid achieves computational efficiency and robust expressiveness, overcoming the limitations of traditional attention-based models. With its superior performance benchmarks and potential for handling increasingly large and complex datasets, Orchid sets a new benchmark for deep learning models in sequence modeling tasks.
Check out the Paper. All credit for this research goes to the researchers of this project. Also, don’t forget to follow us on LinkedIn. Do join our active AI community on Discord.
If you like our work, you will love our Newsletter 📰